 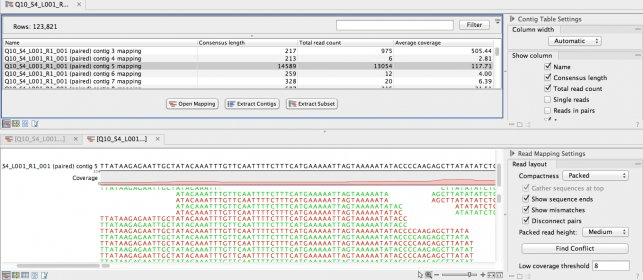
First-time users of CLC Gx on our workstation computers must complete the Workstation Request Form.All USC users can freely access the software on our workstation computers.Equipped with dual-CPU and 512GB RAM, one of our workstation computers is configured specifically to handle large data set and computationally intensive tasks such as de novo genome assembly and sequencing alignment.Wilson Dental Library, the University Park Campus.Norris Medical Library (RM203A), the Health Sciences Campus.The software has been installed on multiple workstation computers:.to the new reference transcriptome using the CLC Genomic Workbench RNAseq. On workstation computers in the libraries General information about installing and upgrading Workbenches. included gene sets of 22 Platyhelminthes species from Wormbase Parasite.Mandatory registration is required for installing CLC Gx on your computer. Please submit the Local Installation Request Form.The computer must be connected to the USC network, either via Ethernet cable on campus or via USC VPN when using wireless (applies to both on- and off-campus wireless connections).Minimum hardware requirement for de novo assembly, metagenomics, and raw reads alignment:: 32GB RAM and Intel i7-6700 or faster processor.Minimum hardware requirement for general use: 16GB RAM and Intel i7-2600 or faster processor.  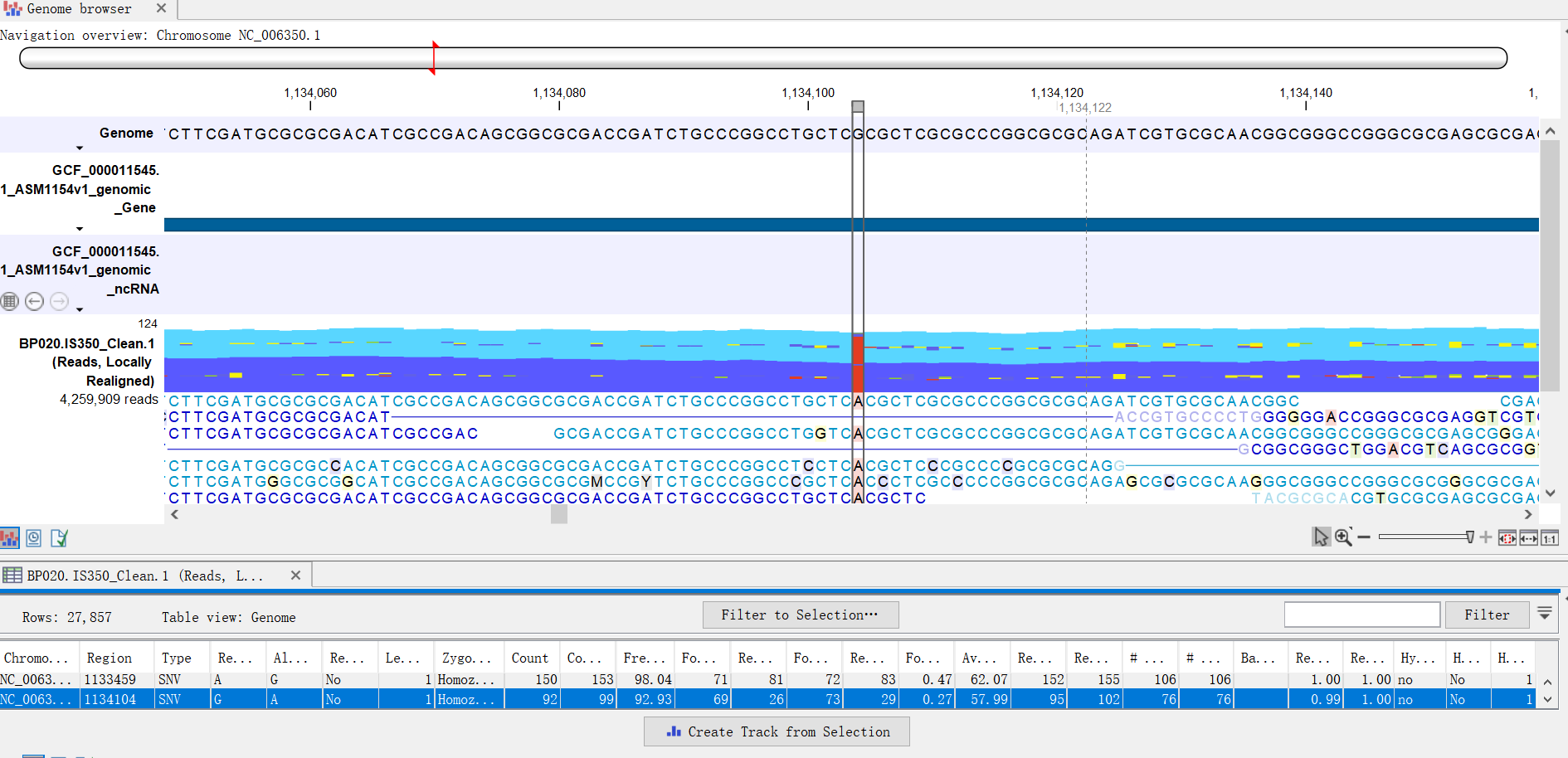
The license consists of TWO concurrent user seats. USC has licensed CLC Gx for the free use of USC faculty, students and staff. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
